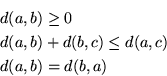
The word ``sparsity distance'' was mentioned above. This is a metric for which I could find no existing name. If one exists I may be confusing things, if not, I invented a metric.
Consider two rows in a sparse matrix. If the two rows are identical, eg. they have zeros and non-zeros at the same column positions, the sparsity distance is 0. If their patterns differ by one non-zero somewhere, the distance is 1, and so on.
The weighted sparsity distance, is a variant of the simple metric from above. The weighted metric gives different weights to pattern differences at different locations. A pattern difference which is located below the diagonal in the system matrix is given a higher weight than a difference located above the diagonal. This is usable in relation to QR factorization, because introducing non-zeros below the diagonal is more expensive than introducing them above the diagonal (below the diagonal will require Givens rotation, above the diagonal requires only back-substitution).
Both distance measures can be shown to be true metrics, eg. they obey the following:
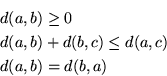
This was a very loose description of the metrics. They are however not important for the understanding of the optimizer, and they are only used in a small isolated part of the program. I felt however, that they deserved to be mentioned.